
This AOP is licensed under the BY-SA license. This license allows reusers to distribute, remix, adapt, and build upon the material in any medium or format, so long as attribution is given to the creator. The license allows for commercial use. If you remix, adapt, or build upon the material, you must license the modified material under identical terms.
AOP: 260
Title
CYP2E1 activation and formation of protein adducts leading to neurodegeneration
Short name
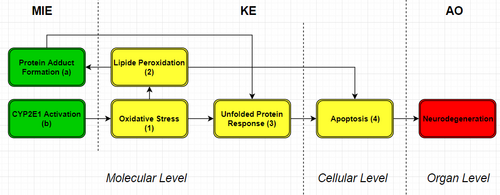
Graphical Representation
Point of Contact
Contributors
- Jelle Broeders
- Marvin Martens
Coaches
OECD Information Table
| OECD Project # | OECD Status | Reviewer's Reports | Journal-format Article | OECD iLibrary Published Version |
|---|---|---|---|---|
This AOP was last modified on July 23, 2024 22:02
Revision dates for related pages
| Page | Revision Date/Time |
|---|---|
| CYP2E1 Activation | April 09, 2018 11:02 |
| Protein Adduct Formation | March 26, 2018 09:45 |
| Lipid Peroxidation | April 04, 2018 14:33 |
| Unfolded Protein Response | September 14, 2023 08:51 |
| General Apoptosis | October 18, 2023 12:20 |
| Neurodegeneration | April 04, 2018 14:53 |
| Increase, Oxidative Stress | February 11, 2026 07:05 |
| CYP2E1 Activation leads to Increase, Oxidative Stress | July 23, 2024 21:56 |
| Increase, Oxidative Stress leads to Lipid Peroxidation | July 23, 2024 21:59 |
| Lipid Peroxidation leads to Protein Adduct Formation | April 05, 2018 04:18 |
| Protein Adduct Formation leads to Unfolded Protein Response | April 05, 2018 04:25 |
| Increase, Oxidative Stress leads to Unfolded Protein Response | July 23, 2024 22:00 |
| Lipid Peroxidation leads to General Apoptosis | April 05, 2018 04:48 |
| Unfolded Protein Response leads to General Apoptosis | April 05, 2018 04:51 |
| General Apoptosis leads to Neurodegeneration | April 05, 2018 04:53 |
| Acetaminophen | November 29, 2016 18:42 |
| Enflurane | April 05, 2018 06:31 |
| Halothane | April 05, 2018 06:32 |
| Isoflurane | April 05, 2018 06:32 |
| Methoxyflurane | April 05, 2018 06:32 |
| Sevoflurane | April 05, 2018 06:33 |
| Chemical:584015 (1-~13~C)Aniline | April 05, 2018 06:33 |
| Chlorzoxazone | April 05, 2018 06:34 |
| Titanium oxide (TiO) | April 05, 2018 06:35 |
| Isoniazid | April 05, 2018 06:36 |
| Ethanol | April 05, 2018 06:38 |
Abstract
The AOP has two different MIEs: protein adduct formation (MIEa) and CYP2E1 activation (MIEb). Protein adduct formation is the interaction between a chemical, or reactive metabolite, and a protein at molecular level. During this interaction a covalent bond is formed which occurs due to the reaction between an electrophilic chemical and the nucleophilic part of a protein. When a chemical forms a covalent bond with a protein the protein is damaged and can loses its function. Acetaldehyde, the metabolite of ethanol, is also one of these chemicals known to form protein adducts. This is why protein adduct formation is added in this AOP based on ethanol. CYP2E1 is one of the enzymes responsible for the metabolism of ethanol, and because of this metabolic activity the MIE in added in this AOP. CYP2E1 participates in the metabolism of endogenous, small and hydrophobic compounds using a oxidation reaction. CYP2E1 is mainly expressed in rat liver cells, but can also be found in rat brain cells. Furthermore, in the human brain CYP2E1 expression is mainly found in the amygdala and prefrontal cortex. At higher concentrations of ethanol the expression of CYP2E1 increases, as well as the activity of CYP2E1 since it has a relatively high Km value for ethanol. In this AOP four different KEs are used, which are oxidative stress (KE1), lipid peroxidation (KE2), unfolded protein response (UPR) (KE3) and apoptosis (KE4). Oxidative stress can be defined as the imbalance between ROS and defence mechanisms against these ROS. ROS levels in a cell can rise which leads to damage by the oxidizing free radicals. Lipid peroxidation is a form of direct damage to lipids in the cell membrane or organelle membranes. The cell membrane will eventually break due to the build-up of all the damage. MDA and 4-hydroxynonenal (HNE) are two products of lipid peroxidation. UPR is a reaction activated by stress in the endoplasmic reticulum (ER). ER stress can be induced by too much protein folding which reaches a higher level than the folding capacity. Also accumulation of unfolded protein in the ER and protein adducts formation with important endoplasmic proteins can induce ER stress, which activates UPR. The final KE is apoptosis, which is programmed cell death in general. The process of apoptosis is well regulated and several signal proteins are known to induce the apoptotic process.
AOP Development Strategy
Context
Strategy
Summary of the AOP
Events:
Molecular Initiating Events (MIE)
Key Events (KE)
Adverse Outcomes (AO)
| Type | Event ID | Title | Short name |
|---|
| MIE | 1508 | CYP2E1 Activation | CYP2E1 Activation |
| MIE | 1509 | Protein Adduct Formation | Protein Adduct Formation |
| KE | 1392 | Increase, Oxidative Stress | Increase, Oxidative Stress |
| KE | 1511 | Lipid Peroxidation | Lipid Peroxidation |
| KE | 1512 | Unfolded Protein Response | Unfolded Protein Response |
| KE | 1513 | General Apoptosis | General Apoptosis |
| AO | 1514 | Neurodegeneration | Neurodegeneration |
Relationships Between Two Key Events (Including MIEs and AOs)
| Title | Adjacency | Evidence | Quantitative Understanding |
|---|
| CYP2E1 Activation leads to Increase, Oxidative Stress | adjacent | Moderate | Moderate |
| Increase, Oxidative Stress leads to Lipid Peroxidation | adjacent | High | High |
| Lipid Peroxidation leads to Protein Adduct Formation | adjacent | High | High |
| Protein Adduct Formation leads to Unfolded Protein Response | adjacent | Moderate | Moderate |
| Increase, Oxidative Stress leads to Unfolded Protein Response | adjacent | Moderate | Moderate |
| Lipid Peroxidation leads to General Apoptosis | adjacent | High | High |
| Unfolded Protein Response leads to General Apoptosis | adjacent | High | High |
| General Apoptosis leads to Neurodegeneration | adjacent | High | High |
Network View
Prototypical Stressors
Life Stage Applicability
Taxonomic Applicability
| Term | Scientific Term | Evidence | Link |
|---|---|---|---|
| human | Homo sapiens | NCBI |
Sex Applicability
Overall Assessment of the AOP
Domain of Applicability
Essentiality of the Key Events
|
Key Event |
Essentiality |
|
MIEa (Protein Adduct Formation) |
Moderate support. Activation of MIEa induces increased activation of KE 3, but direct evidence is not available. One theory about the mechanism is that adducts are formed with critical ER proteins. (Haberzettl, P. & Hill, B. G., 2013; Galligan, J. J. et al., 2014; Cumaoglu, A. et al., 2014; Kessova, I. G. & Cederbaum, A. I., 2005; Huličiak, M. et al., 2012; Sadrieh, N. & Thomas, P. E., 1994; Shin, N. Y. et al., 2007; Sapkota, M. & Wyatt, T. A., 2015; Tuma, D. J., 2002) |
|
MIEb (CYP2E1 Activation) |
High support. Direct evidence is available which prevents the upstream KE 1. CYP2E1 knockout as well as inhibition studies are performed. Activation of CYP2E1 by stressors also showed an increased. (Valencia-Olvera, A. C. et al., 2014; Haorah, J. et al, 2008; Luo, J., 2014; Yang, L. & Cederbaum, A., 2011; Lakshman, M. R. et al., 2013; Jimenez-Lopez, J. M. & Cederbaum, A. I., 2005; Gonzalez, F. J., 2005; Albano, E. et al., 1996; Albano, E., 2006; Wu, D. et al., 2012; Cederbaum, A. I., 2010; Lu, Y. et al., 2010; Oneta, C. M. et al., 2002; Lieber, C. S., 2004; Emerit, J. et al., 2004) |
|
KE 1 (Oxidative Stress) |
High support. Direct and indirect evidence is available for the essentiality of KE 1. Blocking ROS formation inhibits upstream KE 2 and KE 3. The indirect evidence showed that higher ROS induction showed an increased activity of upstream KE 2 and KE 3. |
|
KE 2 (Lipid Peroxidation) |
High support. There is much indirect evidence available showing that inducement of lipid peroxidation can increase activity of MIEb and KE 4. The direct evidence of blocking HNE which results in inhibition of upstream KE 4 shows that there is link between KE 2 and KE 4 and that the underlying molecular pathway is known. |
|
KE 3 (UPR) |
Moderate support. There is direct as well as indirect evidence available which shows molecular understanding of how KE 3 can induce KE 4. The uncertainty lies in whether ER stress alone can induce KE 4, or that KE 1 also plays a role in it. This is more discussed in detail in chapter 4. |
|
KE 4 (Apoptosis) |
High support. Neurodegeneration is the loss of neuron cells in the brain. |
See table below where an overview is provided of the direct and indirect evidence. For the meaning of numbering see Abstract and the image of the AOP,
|
Key event relatio-nship |
Methods |
Influence on downstream Key events |
Direct/Indirect evidence |
|
KER 1: MIEb ---> KE1 |
1. Stimulation of CYP2E1 by stressors in rat livers. 2. Inhibition studies of CYP2E1 in neuron cells. 3. CYP2E1 KO in mice where TBARS values are measured. 4. Induction of CYP2E1 results in higher ROS levels an higher CYP2E1 expression, study performed in granule neuron cells. |
1. Activation KE 1 2. Inhibition KE 1 3. Inhibition KE 1 4. Activation KE 1 |
1. Indirect 2. Direct 3. Direct 4. Indirect |
|
KER 2: KE1 ---> KE2 |
1. Lower ROS level by adding higher concentrations of antioxidants or resveratrol (inhibitor of ROS). TBARS and LOOH product was measured in rat microsomes. 2. Correlation study where higher ROS levels increased lipid peroxidation in aging brains. |
1. Inhibition KE 2 2. Activation KE 2 |
1. Direct 2. Indirect |
|
KER 3: KE1 ---> KE3 |
1. Lower ROS levels by overexpression of antioxidant SOD1, NAC or GSH resulted in induction of UPR markers. Measured in neuron cells. 2. Stimulation of ROS formation by ethanol, which induces the UPR response in 2 hours after exposure. |
1. Inhibition KE 3 2. Activation KE 3 |
1. Direct 2. Indirect |
|
KER 4: KE2 ---> MIEa |
1.Proteomic detection techniques for HNE adducts, HNE is a reactive aldehyde product of lipid peroxidation. 2. SERS monitoring detection, showed link between increased lipid peroxidation and increased protein adduct formation. |
1. Activation MIEb 2. Activation MIEb |
1. Indirect 2. Indirect |
|
KER 5: MIEa ---> KE3 |
1. HNE (known to form protein adducts) treatment in rat aortic smooth muscle cells induced expression of the PERK pathway, which is part of the UPR. Same study is also performed in different settings. 2. Some toxicants can form protein adducts with ER proteins, what can induce ER stress and the UPR. |
1. Activation KE 3 2. Activation KE 3 |
1. Indirect 2. Indirect |
|
KER 6: KE2 ---> KE4 |
1. HNE can induce Fas/CD95DR expression, which regulated the extrinsic pathway of apoptosis. 2. Knockout of GSTA4 in mouse, which is an antioxidant for HNE, showed an increase in Fas expression. 3. ASK1 and JNK are activated by Fas. Increased HNE concentrations showed higher expression of ASK1 and JNK. When Fas was inhibited apoptosis was stopped. 4. HNE induces mitochondrial dysfunction which leads to apoptosis. Higher HNE levels showed increased expression of cytochrome c and caspases. Caspase 3 and 9 are mainly activated. Both are part of the intrinsic pathway of apoptosis. |
1. Activation KE 4 2. Inhibition KE 4 3. Activation KE 4 4. Activation KE 4 |
1. Indirect 2. Direct 3. Indirect 4. Indirect |
|
KER 7: KE3 ---> KE4 |
1. Higher expression of IRE1 and PERK, which are UPR markers, showed an increase of caspases expression. These caspases play a major role in the apoptotic pathway. 2. Inhibition of ER stress by DHCR24 resulted in a lower level of CHOP expression. Also an inhibition of apoptosis was shown. |
1. Activation KE 4 2. Inhibition KE 4 |
1. Indirect 2. Direct |
|
KER 8: KE4 ---> AO |
1. Neuron loss is detected in neurodegenerative diseases, such as Alzheimer. |
1. Activation AO |
1. Indirect |
Evidence Assessment
Known Modulating Factors
Quantitative Understanding
In AOP1 there are some knowledge gaps present which is one of the principles of the AOP concept. CYP2E1 activation is known to increase the ROS concentration in a cell, but the underlying mechanism is not completely understood. There are two main mechanisms which are suggested in literature, either CYP2E1 or NADPH oxidase could be the primary enzyme which is responsible for ROS formation and cause the further damage in the cells. NAPDH oxidase recycles the NADP+ which is formed during the reaction cycle of CYP2E1, during this cycle ROS is formed due to the uncoupling reaction. CYP2E1 shows a relatively high activity of NADPH oxidase activity and is poorly coupled with NADPH-cytochrome P450 reductase. When NADPH oxidase is inhibited by anti-CYP2E1 IgG a reduction of ROS induced lipid peroxidation was shown. Knock-out or inhibition of CYP2E1 itself resulted in lower oxidative stress. A study performed by Bradford et al. showed that NADPH oxidase knock-out mice attenuated liver injury, where CYP2E1 knock-out mice did not show attenuating of liver injury. On the other hand, NADHP oxidase knock-out mice did not reduce oxidative stress damage to DNA, where CYP2E1 knock-out mice did reduced the damage. Another study by Zhang et al. looked at the influence of NADPH oxidase, an inhibiter against NADPH oxidase was used which reduced the formation of ROS in PC12 cells. Finally, Shah et al. and Furukawa et al. also showed that NADPH oxidase inhibition leads to a reduced formation of ROS, both studies were done in different disease context. The principle of ROS formation by NADPH oxidase is the formation of H2O2 since O2 is used as a substrate. By the Fenton-Weiss-Haber reaction multiple oxidants can be produces. But as mentioned above, several studies showed that CYP2E1 inhibition alone is enough to reduce ROS formation. To take into account, studies described above are all done in liver cells. The mechanism of CYP2E1 activation could be different in the brain.
Another knowledge gap is the mechanism of protein adducts that can induce ER stress, and ultimately the UPR. The assumed mechanism is that protein adducts are formed with critical ER proteins, which leads to the dysfunction of the ER. Furthermore, it is also a possibility that protein adducts inhibit the folding of proteins. These proteins can accumulate in the ER and when the protein accumulation is higher than the capacity ER stress is induced. Further research must be done to define the mechanism of how ER stress is induced by protein adducts, which will eventually lead to the UPR.
Considerations for Potential Applications of the AOP (optional)
References
Leist, M. et al. Adverse outcome pathways: opportunities, limitations and open questions. Arch. Toxicol. 204, 1–29 (2017).
LoPachin, R. M. & DeCaprio, A. P. Protein adduct formation as a molecular mechanism in neurotoxicity. Toxicological Sciences 86, 214–225 (2005).
Cederbaum, A. I. Alcohol Metabolism. Clinics in Liver Disease 16, 667–685 (2012).
Sapkota, M. & Wyatt, T. A. Alcohol, aldehydes, adducts and airways. Biomolecules 5, 2987–3008 (2015).
Tuma, D. J. Role of malondialdehyde-acetaldehyde adducts in liver injury. Free Radical Biology and Medicine 32, 303–308 (2002).
Neafsey, P. et al. Genetic polymorphism in CYP2E1: Population distribution of CYP2E1 activity. Journal of Toxicology and Environmental Health - Part B: Critical Reviews 12, 362–388 (2009).
Zimatkin, S. M., Pronko, S. P., Vasiliou, V., Gonzalez, F. J. & Deitrich, R. A. Enzymatic mechanisms of ethanol oxidation in the brain. Alcohol. Clin. Exp. Res. 30, 1500–1505 (2006).
Toselli, F. et al. Expression of CYP2E1 and CYP2U1 proteins in amygdala and prefrontal cortex: Influence of alcoholism and smoking. Alcohol. Clin. Exp. Res. 39, 790–797 (2015).
Zakhari, S. Alcohol metabolism and epigenetics changes. Alcohol Res. 35, 6–16 (2013).
Valencia-Olvera, A. C., Morán, J., Camacho-Carranza, R., Prospéro-García, O. & Espinosa-Aguirre, J. J. CYP2E1 induction leads to oxidative stress and cytotoxicity in glutathione-depleted cerebellar granule neurons. Toxicol. Vitr. 28, 1206–1214 (2014).
Ayala, A., Muñoz, M. F. & Argüelles, S. Lipid peroxidation: Production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxidative Medicine and Cellular Longevity 2014, (2014).
Foufelle, F. & Fromenty, B. Role of endoplasmic reticulum stress in drug-induced toxicity. Pharmacol. Res. Perspect. 4, e00211 (2016).
Shtilbans, V., Wu, M. & Burstein, D. E. Evaluation of apoptosis in cytologic specimens. Diagnostic Cytopathology 38, 685–697 (2010).
Wu, J., Sun, J. & Xue, Y. Involvement of JNK and P53 activation in G2/M cell cycle arrest and apoptosis induced by titanium dioxide nanoparticles in neuron cells. Toxicol. Lett. 199, 269–276 (2010).
Redza-Dutordoir, M. & Averill-Bates, D. A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta - Mol. Cell Res. 1863, 2977–2992 (2016).
Cederbaum, A. I. Role of Cytochrome P450 and Oxidative Stress in Alcohol-Induced Liver Injury. AIMSCI Inc. (2017).
Haorah, J. et al. Mechanism of alcohol-induced oxidative stress and neuronal injury. Free Radic. Biol. Med. 45, 1542–1550 (2008).
Sapkota, M. & Wyatt, T. A. Alcohol, aldehydes, adducts and airways. Biomolecules 5, 2987–3008 (2015).
Tuma, D. J. Role of malondialdehyde-acetaldehyde adducts in liver injury. Free Radical Biology and Medicine 32, 303–308 (2002).
Valencia-Olvera, A. C., Morán, J., Camacho-Carranza, R., Prospéro-García, O. & Espinosa-Aguirre, J. J. CYP2E1 induction leads to oxidative stress and cytotoxicity in glutathione-depleted cerebellar granule neurons. Toxicol. Vitr. 28, 1206–1214 (2014).
Ayala, A., Muñoz, M. F. & Argüelles, S. Lipid peroxidation: Production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxidative Medicine and Cellular Longevity 2014, (2014).
Uttara, B., Singh, A. V, Zamboni, P. & Mahajan, R. T. Oxidative stress and neurodegenerative diseases: a review of upstream and downstream antioxidant therapeutic options. Curr. Neuropharmacol. 7, 65–74 (2009).
Andringa, K. K., Udoh, U. S., Landar, A. & Bailey, S. M. Proteomic analysis of 4-hydroxynonenal (4-HNE) modified proteins in liver mitochondria from chronic ethanol-fed rats. Redox Biol. 2, 1038–1047 (2014).
Albano, E. et al. Role of cytochrome P4502E1-dependent formation of hydroxyethyl free radical in the development of liver damage in rats intragastrically fed with ethanol. Hepatology 23, 155–163 (1996).
Wu, D., Wang, X., Zhou, R., Yang, L. & Cederbaum, A. I. Alcohol steatosis and cytotoxicity: The role of cytochrome P4502E1 and autophagy. Free Radic. Biol. Med. 53, 1346–1357 (2012).
Lu, Y., Wu, D., Wang, X., Ward, S. C. & Cederbaum, A. I. Chronic alcohol-induced liver injury and oxidant stress are decreased in cytochrome P4502E1 knockout mice and restored in humanized cytochrome P4502E1 knock-in mice. Free Radic. Biol. Med. 49, 1406–1416 (2010).
Sultana, R., Perluigi, M. & Butterfield, D. A. Lipid peroxidation triggers neurodegeneration: A redox proteomics view into the Alzheimer disease brain. Free Radical Biology and Medicine 62, 157–169 (2013).
Devasagayam, T. P. A., Kamat, J. P., Mohan, H. & Kesavan, P. C. Caffeine as an antioxidant: Inhibition of lipid peroxidation induced by reactive oxygen species. Biochim. Biophys. Acta - Biomembr. 1282, 63–70 (1996).
Leutner, S., Eckert, A. & Müller, W. E. ROS generation, lipid peroxidation and antioxidant enzyme activities in the aging brain. J. Neural Transm. 108, 955–967 (2001).
Nosáľ, R. et al. On the molecular pharmacology of Resveratrol on oxidative burst inhibition in professional phagocytes. Oxid. Med. Cell. Longev. 2014, (2014).
Ramirez-Alvarado, M., Kelly, J. W. & Dobson, C. M. Protein Misfolding Diseases: Current and Emerging Principles and Therapies. Protein Misfolding Diseases: Current and Emerging Principles and Therapies (2010). doi:10.1002/9780470572702
Hayashi, T. et al. Damage to the endoplasmic reticulum and activation of apoptotic machinery by oxidative stress in ischemic neurons. J. Cereb. Blood Flow Metab. 25, 41–53 (2005).
Chen, G. et al. Ethanol promotes endoplasmic reticulum stress-induced neuronal death: Involvement of oxidative stress. J. Neurosci. Res. 86, 937–946 (2008).
Tsedensodnom, O., Vacaru, A. M., Howarth, D. L., Yin, C. & Sadler, K. C. Ethanol metabolism and oxidative stress are required for unfolded protein response activation and steatosis in zebrafish with alcoholic liver disease. Dis. Model. Mech. 6, 1213–1226 (2013).
Cano, M. et al. Oxidative stress induces mitochondrial dysfunction and a protective unfolded protein response in RPE cells. Free Radic. Biol. Med. 69, 1–14 (2014).
Poli, G. et al. Enzymatic impairment induced by biological aldehydes in intact rat liver cells. Res. Commun. Chem. Pathol. Pharmacol. 38, (1982).
Siems, W. & Grune, T. Intracellular metabolism of 4-hydroxynonenal. in Molecular Aspects of Medicine 24, 167–175 (2003).
Codreanu, S. G., Zhang, B., Sobecki, S. M., Billheimer, D. D. & Liebler, D. C. Global analysis of protein damage by the lipid electrophile 4-hydroxy-2-nonenal. Mol. Cell. Proteomics 8, 670–80 (2009).
Gong, T. et al. Rapid SERS monitoring of lipid-peroxidation-derived protein modifications in cells using photonic crystal fiber sensor. Journal of Biophotonics 9, 32–37 (2016).
Haberzettl, P. & Hill, B. G. Oxidized lipids activate autophagy in a JNK-dependent manner by stimulating the endoplasmic reticulum stress response. Redox Biol. 1, 56–64 (2013).
Galligan, J. J. et al. Oxidative stress-mediated aldehyde adduction of GRP78 in a mouse model of alcoholic liver disease: Functional independence of ATPase activity and chaperone function. Free Radic. Biol. Med. 73, 411–420 (2014).
Cumaoglu, A., Arıcıoglu, A. & Karasu, C. Redox status related activation of endoplasmic reticulum stress and apoptosis caused by 4-hydroxynonenal exposure in INS-1 cells. Toxicol. Mech. Methods 24, 362–367 (2014).
Kessova, I. G. & Cederbaum, A. I. The effect of CYP2E1-dependent oxidant stress on activity of proteasomes in HepG2 cells. J Pharmacol Exp Ther 315, 304–312 (2005).
Huličiak, M. et al. Covalent binding of cisplatin impairs the function of Na +/K +-ATPase by binding to its cytoplasmic part. Biochem. Pharmacol. 83, 1507–1513 (2012).
Sadrieh, N. & Thomas, P. E. Characterization of rat cytochrome P450 isozymes involved in the covalent binding of cyclosporin A to microsomal proteins. Toxicol. Appl. Pharmacol. 127, 222–232 (1994).
Shin, N. Y., Liu, Q., Stamer, S. L. & Liebler, D. C. Protein targets of reactive electrophiles in human liver microsomes. Chem. Res. Toxicol. 20, 859–867 (2007).
Dalleau, S., Baradat, M., Guéraud, F. & Huc, L. Cell death and diseases related to oxidative stress: 4-hydroxynonenal (HNE) in the balance. Cell Death Differ. 20, 1615–30 (2013).
Li, J. et al. Regulation of CD95 (Fas) expression and Fas-mediated apoptotic signaling in HLE B-3 cells by 4-hydroxynonenal. Biochemistry 45, 12253–12264 (2006).
Engle, M. R. et al. Physiological role of mGSTA4-4, a glutathione S-transferase metabolizing 4-hydroxynonenal: Generation and analysis of mGsta4 null mouse. Toxicol. Appl. Pharmacol. 194, 296–308 (2004).
Sharma, R. et al. 4-Hydroxynonenal self-limits Fas-mediated DISC-independent apoptosis by promoting export of Daxx from the nucleus to the cytosol and its binding to Fas. Biochemistry 47, 143–156 (2008).
Liu, W., Porter, N. A., Schneider, C., Brash, A. R. & Yin, H. Formation of 4-hydroxynonenal from cardiolipin oxidation: Intramolecular peroxyl radical addition and decomposition. Free Radic. Biol. Med. 50, 166–178 (2011).
Moreira, P. I. et al. Mitochondria: A therapeutic target in neurodegeneration. Biochimica et Biophysica Acta - Molecular Basis of Disease 1802, 212–220 (2010).
Liu, W. et al. 4-Hydroxynonenal Induces a Cellular Redox Status-Related Activation of the Caspase Cascade for Apoptotic Cell Death. J. Cell Sci. 113 ( Pt 4, 635–641 (2000).
Knoll, N. et al. Genotoxicity of 4-hydroxy-2-nonenal in human colon tumor cells is associated with cellular levels of glutathione and the modulation of glutathione S-transferase A4 expression by butyrate. Toxicol. Sci. 86, 27–35 (2005).
Hiramatsu, N. et al. Translational and posttranslational regulation of XIAP by eIF2 and ATF4 promotes ER stress-induced cell death during the unfolded protein response. Mol. Biol. Cell 25, 1411–1420 (2014).
Shah, A. & Kumar, A. Methamphetamine-mediated endoplasmic reticulum (ER) stress induces type-1 programmed cell death in astrocytes via ATF6, IRE1 beta and PERK pathways. Oncotarget 7, 46100–46119 (2016).
Han, J. et al. ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat. Cell Biol. 15, 481–490 (2013).
Lu, X. et al. 3 ??-hydroxysteroid-?? 24 reductase (DHCR24) protects neuronal cells from apoptotic cell death induced by Endoplasmic Reticulum (ER) stress. PLoS One 9, (2014).
Shah, A., Kumar, S., Simon, S. D., Singh, D. P. & Kumar, A. HIV gp120- and methamphetamine-mediated oxidative stress induces astrocyte apoptosis via cytochrome P450 2E1. Cell Death Dis. 4, (2013).
Ekström, G. & Ingelman-Sundberg, M. Rat liver microsomal NADPH-supported oxidase activity and lipid peroxidation dependent on ethanol-inducible cytochrome P-450 (P-450IIE1). Biochem. Pharmacol. 38, 1313–1319 (1989).
Lu, Y. & Cederbaum, A. I. CYP2E1 and oxidative liver injury by alcohol. Free Radical Biology and Medicine 44, 723–738 (2008).
Rashba-Step, J., Turro, N. J. & Cederbaum, A. I. Increased NADPH- and NADH-dependent production of superoxide and hydroxyl radical by microsomes after chronic ethanol treatment. Arch Biochem Biophys 300, 401–408 (1993).
Bradford, B. U. et al. Cytochrome P450 CYP2E1, but not nicotinamide adenine dinucleotide phosphate oxidase, is required for ethanol-induced oxidative DNA damage in rodent liver. Hepatology 41, 336–344 (2005).
Furukawa, S. et al. Increased oxidative stress in obesity and its impact on metabolic syndrome. J. Clin. Invest. 114, 1752–1761 (2004).