
This AOP is licensed under the BY-SA license. This license allows reusers to distribute, remix, adapt, and build upon the material in any medium or format, so long as attribution is given to the creator. The license allows for commercial use. If you remix, adapt, or build upon the material, you must license the modified material under identical terms.
AOP: 46
Title
AFB1: Mutagenic Mode-of-Action leading to Hepatocellular Carcinoma (HCC)
Short name
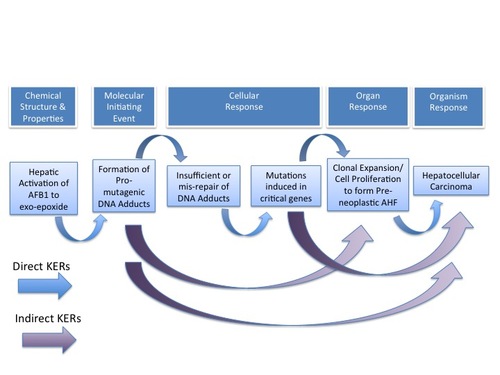
Graphical Representation
Point of Contact
Contributors
- Ted Simon
Coaches
OECD Information Table
| OECD Project # | OECD Status | Reviewer's Reports | Journal-format Article | OECD iLibrary Published Version |
|---|---|---|---|---|
| 1.8 | Under Review |
This AOP was last modified on April 29, 2023 16:02
Revision dates for related pages
| Page | Revision Date/Time |
|---|---|
| Formation, Pro-mutagenic DNA Adducts | September 16, 2017 10:15 |
| Tumorigenesis, Hepatocellular carcinoma | November 29, 2016 19:08 |
| Increased, Induced Mutations in Critical Genes | September 16, 2017 10:15 |
| Metabolism of AFB1, Production of Reactive Electrophiles | September 16, 2017 10:15 |
| Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) | September 16, 2017 10:15 |
| Increased, Insufficient repair or mis-repair of pro-mutagenic DNA adducts | September 16, 2017 10:15 |
| Increased, Induced Mutations in Critical Genes leads to Tumorigenesis, Hepatocellular carcinoma | November 30, 2016 14:12 |
| Increased, Induced Mutations in Critical Genes leads to Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) | November 29, 2016 20:19 |
| Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) leads to Tumorigenesis, Hepatocellular carcinoma | November 29, 2016 20:19 |
| Increased, Insufficient repair or mis-repair of pro-mutagenic DNA adducts leads to Increased, Induced Mutations in Critical Genes | November 29, 2016 20:19 |
| Formation, Pro-mutagenic DNA Adducts leads to Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) | November 29, 2016 20:19 |
| Formation, Pro-mutagenic DNA Adducts leads to Tumorigenesis, Hepatocellular carcinoma | November 29, 2016 20:06 |
| Metabolism of AFB1, Production of Reactive Electrophiles leads to Formation, Pro-mutagenic DNA Adducts | November 29, 2016 20:10 |
| Formation, Pro-mutagenic DNA Adducts leads to Increased, Insufficient repair or mis-repair of pro-mutagenic DNA adducts | December 03, 2016 16:37 |
Abstract
Aflatoxin B1 (AFB1), with ubiquitous exposure and a rich database, was selected as a case study for development of an AOP on mutagenic MOA for cancer. AFB1 has been determined to induce the AO hepatocellular carcinoma (HCC) via a DNA-reactive MOA in many species, including humans. The sequential KEs identified for AFB1 are as follows: pre-MIE: Hepatic metabolic activation; MIE: Formation of a pro-mutagenic DNA adduct (N7-AFB1-guanine or AFB1-FAPy); KE#1: Inadequate or mis-repair of the pro-mutagenic DNA adducts; KE#2: Induced mutation in critical gene(s); KE#3: Cellular proliferation and clonal expansion of mutant cells (pre-neoplastic lesions); AO: HCC
AOP Development Strategy
Context
This AOP describes a mutagenic mode of action (MOA) for cancer, using Aflatoxin B1 (AFB1) as a case example. Mutagenic MOAs are distinguished from other MOAs for cancer in that the chemical induces mutations in genes that are involved in the etiology of the cancer. This MOA is distinguished from a non-mutagenic MOA for carcinogens that cause proliferation of cells with existing mutations, or that in some other way promote the growth of cancer gene mutant cells. The induction of mutation in the cancer critical gene is the key event (KE) that is unique to mutagenic MOA (Jarabek et al., 2009; Pottenger et al., 2014). In this AFB1 case example, mutation in critical genes is a consequence of formation of pro-mutagenic DNA adducts by metabolites of the chemical. It is important to note that all cancers involve an increase in cells containing mutations in cancer critical gene(s). Likewise, cell proliferation is an essential step in the formation of tumors. To establish that an agent has a mutagenic MOA, it is necessary to determine the key events, both in terms of temporality and of dose-response concordance, between the increase in the number of mutant cells, cell proliferation, the appearance of any pre-neoplastic lesions, and ultimately tumor occurrence. The types of data that can be used to investigate whether a chemical acts via a mutagenic MOA include information on the chemical’s ability to cause mutations, the temporality of those induced mutations, and the type of mutations that the chemical induces. It is not sufficient to determine if the chemical can induce mutations in any one of a number of standard gene mutation assays. Furthermore, because the etiology of all tumors includes an increase in cells with cancer gene mutations, the presence of mutations in the tumor tissue does not provide definitive information on MOA. When there is a high frequency of tumors with specific mutations (as is the case for AFB1), this observation provides a hypothesis that can be further evaluated.
The most definitive level of proof that a chemical acts via a mutagenic MOA is the demonstration that the chemical can induce the specific cancer gene mutation(s) observed in a majority of the specific tumors, and that the formation of this mutation is an early KE in the AOP (Moore et al., 2008). In the absence of this information, the ability of the chemical to induce the type(s) of mutations seen in the majority of the specific tumors adds greatly to the weight of evidence. Such information on specific chemical-induced mutations in cancer critical genes is uncommon, and currently, no such information is available for AFB1. A newly developed method, allele specific competitive blocker-polymerase chain reaction (ACB-PCR) has proved useful in providing such information, and data on specific chemical-induced mutations are available for only a very small number of chemicals (Parsons et al., 2010).
Both in vitro and in vivo gene mutation assays can detect chemically induced mutations; by sequencing the DNA from mutants induced in these assays, the type(s) of mutations that a chemical induces can be determined. A match between the type(s) of mutations found in these surrogate gene mutation assays and the mutations seen in a majority of the tumors provides a high level of confidence that the chemical has a mutagenic MOA for the induction of those specific tumors. Fortunately this type of data is available for AFB1.
This AOP is written for AFB1, a data-rich chemical. It is clear that (1) AFB1 can induce mutations in gene mutation assays; (2) AFB1 induces hepatocellular carcinoma (HCC) in a variety of species, including humans; (3) there is a high frequency of a specific cancer gene mutation (codon 249 of p53) in the human HCCs found in people in regions with high AFB1 exposure; and (4) the type of mutation seen in the human tumor (codon 249 of p53) is the same type of mutation that is seen in the surrogate gene mutation assay. Thus there is a high level of confidence that AFB1 has a mutagenic MOA for HCC in humans.
The AOP for a mutagenic MOA for liver cancer induced by Aflatoxin B1 includes the following series of key events: • pre-MIE: Hepatic metabolic activation • MIE: Pro-mutagenic DNA adduct formation • KE#1: Insufficient repair or mis-repair of pro-mutagenic DNA adducts • KE#2: Induced mutation in critical genes • KE#3: Cellular proliferation and clonal expansion of mutant cells (pre-neoplastic lesions) • AO: Hepatocellular carcinoma (HCC)
This AOP describes the KE that are specific to a mutagenic MOA for AFB1 (e.g., [pre-Molecular Initiating Event or pre-MIE: metabolic activation,] MIE: pro-mutagenic AFB1 DNA adduct, KE#1: insufficient/mis-repair of DNA, KE#2: induced mutation in critical gene). In addition, this AOP includes all of the steps involved from the exposure to AFB1 through to the appearance of the HCC, the adverse outcome. KE#3, the occurrence and growth of altered hepatic foci and the subsequent development of HCC (as described below) are events that would be involved in the etiology of all HCC whether occurring through a mutagenic MOA or another, different MOA. Although AFB1-specific information is lacking for these later steps in the AOP, the accumulated scientific knowledge of the development of HCC includes the recognition that all AOPs leading to HCC have a number of general steps in common; thus, these general steps are included in this AOP and are identified as not being specific to AFB1-induced HCC.
Strategy
Summary of the AOP
Events:
Molecular Initiating Events (MIE)
Key Events (KE)
Adverse Outcomes (AO)
| Type | Event ID | Title | Short name |
|---|
| MIE | 373 | Formation, Pro-mutagenic DNA Adducts | Formation, Pro-mutagenic DNA Adducts |
| KE | 376 | Increased, Induced Mutations in Critical Genes | Increased, Induced Mutations in Critical Genes |
| KE | 409 | Metabolism of AFB1, Production of Reactive Electrophiles | Metabolism of AFB1, Production of Reactive Electrophiles |
| KE | 491 | Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) | Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) |
| KE | 493 | Increased, Insufficient repair or mis-repair of pro-mutagenic DNA adducts | Increased, Insufficient repair or mis-repair of pro-mutagenic DNA adducts |
| AO | 378 | Tumorigenesis, Hepatocellular carcinoma | Tumorigenesis, Hepatocellular carcinoma |
Relationships Between Two Key Events (Including MIEs and AOs)
| Title | Adjacency | Evidence | Quantitative Understanding |
|---|
| Increased, Induced Mutations in Critical Genes leads to Tumorigenesis, Hepatocellular carcinoma | non-adjacent | Moderate | Low |
| Formation, Pro-mutagenic DNA Adducts leads to Clonal Expansion/Cell Proliferation, to form Altered Hepatic Foci (AHF) | non-adjacent | High | High |
| Formation, Pro-mutagenic DNA Adducts leads to Tumorigenesis, Hepatocellular carcinoma | non-adjacent | Moderate | Moderate |
Network View
Prototypical Stressors
Life Stage Applicability
| Life stage | Evidence |
|---|---|
| During development and at adulthood | Moderate |
Taxonomic Applicability
| Term | Scientific Term | Evidence | Link |
|---|---|---|---|
| humans | Homo sapiens | High | NCBI |
| rat | Rattus norvegicus | High | NCBI |
| tree shrew | Tupaia glis | High | NCBI |
| rainbow trout | Oncorhynchus mykiss | High | NCBI |
| guinea pig | Cavia porcellus | High | NCBI |
| Syrian hamsters | Mesocricetus auratus | High | NCBI |
| chickens | Gallus gallus | Moderate | NCBI |
| turkey | Meleagris gallopavo | Moderate | NCBI |
| domestic cattle | Bos taurus | Moderate | NCBI |
| swine | Sus scrofa | Moderate | NCBI |
| dogs | Canis lupus familiaris | Moderate | NCBI |
| cats | Felis catus | Moderate | NCBI |
| rhesus monkeys | Macaca mulatta | Moderate | NCBI |
Sex Applicability
| Sex | Evidence |
|---|---|
| Unspecific | High |
Overall Assessment of the AOP
Domain of Applicability
This AOP is applicable to all life stages, to many vertebrate taxons, and to both genders.
Essentiality of the Key Events
Support for Essentiality of KEs
| Support for Essentiality of KEs | Defining Question | High (Strong) | Moderate | Weak |
| Are downstream key events and/or the AO prevented if an upstream key event is blocked? [e.g., stop/reversibility studies, antagonism, knock out models, etc.) | Multiple lines of experimental evidence illustrating essentiality for several of the key events | There is at least one line of experimental evidence indicating essentiality of an important key event | Indirect or no experimental evidence of the essentiality of any of the key events | |
| Hepatic metabolism | Essentiality of the pre-MIE is Strong.
Rationale: S9 activation, Oltipraz or CDDO-Im chemointervention. Quantity of exo-epoxide formed influences the number of adducts formed (e.g., differences between rats and mice) |
|||
| Formation of Pro-mutagenic DNA adducts | Essentiality of the MIE is Strong.
Rationale: Limited direct empirical support: when total adducts reduced due to oltipraz or CDDO-Im, few or no tumors, which allows inference of the contrary—when adducts present, can get tumors |
|||
| Insufficient or Mis-repair of pro-mutagenic DNA adducts. | Essentiality of the KE1 is Strong
Rationale: Necessary; data from repair deficient yeast demonstrate that mutations are increased when repair is deficient |
|||
| Induced mutations in critical genes | Essentiality of the KE2 is Strong
Rationale: Necessary. While there is no direct data for AFB1 that mutations are “induced” in critical genes, it is clear that having an increased number of cells with mutations in critical genes is required for the altered phenotype that leads to cell proliferation and ultimately to tumors. |
|||
| Clonal Expansion / Cell Proliferation (AHF) | Essentiality of the KE3 is Strong
Rationale: Necessary, chemoprevention studies, HBV and possible effect on proliferation The plethora of longer term initiation-promotion studies provide much evidence for the clonal expansion of foci. |
|||
Chemoprotection
A number of substances have been shown to act as chemoprotectant or chemopreventive agents against AFB1-induced liver tumors [1-3]. These compounds may act at several points in the pathway beginning with exposure to AFB1 and ending with the occurrence of tumors; such data address the essentiality of key events. Below we consider a framework for the action of chemoprotectants based on the KEs in the AOP and then attempt a classification of various chemoprotectants based on this framework with consideration of cellular signaling pathways that likely underlie their chemoprotective action [4,5]. Framework for Chemoprotectant Action
Chemoprotective agents may act in the three distinct ways: • protection against the formation of pro-mutagenic adducts • protection against mutations in tumor-critical locations • protection against the formation of pre-neoplastic lesions and tumors through appropriate cell cycle checkpoints or cytotoxicity
First, chemoprotectants could enhance detoxification metabolic pathways (e.g., GST conjugation) for AFB1 or decrease formation of the pro-mutagenic exo-epoxide metabolite. Second, AFB1 pro-mutagenic adducts may form in locations that lead to critical (pro-tumorigenic) mutations or in locations that are much less likely to lead to critical (pro-tumorigenic) mutations. Chemoprotectants could decrease the likelihood of adducts occurring in tumor-critical locations or enhance the fidelity and efficacy of DNA repair processes, thus reducing the effective numbers and impact of any pro-mutagenic adducts.
Third, chemoprotectants may act to block or lessen the progression from AFB1-induced critical mutations to pre-neoplastic lesions and tumors. Alternatively, chemoprotectants may enhance the ability of cell cycle checkpoint systems to interfere with cell replication (critical to render a mutation permanent and thus heritable), or may produce apoptotic cell death. In addition, chemoprotectants may produce cytotoxicity in cells with a sufficient number of adducts such that the cells die before any mutations can be fixed at the next cell division. Induction of cellular defense against electrophilic and oxidative stress is mediated through a signaling pathway via the transcription factor called nuclear factor erythroid 2-related factor 2 (Nrf2) and the Kelch-like erythroid cell-derived protein 1 (Keap1). Nrf2 turns over rapidly within cells via ubiquitination unless critical cysteine thiols of these two molecules are modified by oxidants or electrophiles. Reduced ubiquitination of Nrf2-Keap1 will increase cellular levels of free Nrf2 and lead to binding of Nrf2 dimers to anti-oxidant response elements (AREs) on DNA, resulting in the induction of anti-oxidant genes. Anti-oxidants such as the food additive butylated hydroxyanisole (BHA), phytochemicals such as genistein, quercetin and curcumin, redox-active bisphenols and quinones, thiocarbamates, dithiolethiones, polyphenols and triterpenoids are all capable of modifying the cysteine residues on Nrf2 and Keap1, and thus they can all serve as ARE inducers.
Nrf2 can induce a large number of ARE-dependent genes including the mixed function oxidase CYP2A5, the Phase 2 enzymes UGT1A1 and SULT3A1, superoxide dismutase, peroxiredoxin, glutathione peroxidase, heme oxygenase 1 (HO-1) and others [6]. In fact, the constellation of ARE-dependent enzymes likely act in all of the ways discussed in the framework above.
The majority of chemoprotective agents appear to work via activation of Keap1 due to cysteine modification.
Actions of Some Specific Chemoprotectants
In this section, the actions of specific chemoprotectants will be considered.
Selenium
Dietary selenium reduced the volume fraction of altered hepatic foci (AHF) occurring in the livers of rats fed a high fat diet ten weeks after initiation with 10 days of intragastric administration of 4 mg/kg AFB1. In rats fed diets containing 0.15 ppm Se or more, glutathione peroxidase was elevated compared with animals fed diets containing <0.02 ppm Se [7]. Se may modify cysteine residues on Nrf2-Keap1.
Oltipraz
The dithiolthione oltipraz was administered to rats at 0.01 to 0.1% in the diet for 1 week prior to administration of 250 μg/kg AFB1 by gavage 5 days per week for two weeks. The rats continued a diet containing oltipraz for one week following cessation of AFB1 administration. The animals were then restored to a control diet and sacrificed twelve weeks later. Oltipraz increased GST activity and reduced the volume of GGT-positive foci in a dose-dependent manner, but did not eliminate their formation completely. In rats administered a single gavage dose of 250 μg/kg (14C)-AFB1 and sacrificed two hours later, binding of the labeled AFB1 to DNA was reduced in a dose-dependent fashion, thus AFB1-induced DNA adduct formation is reduced, probably due to the induction of GST activity. In rats maintained on diets containing 0.01 and 0.1% oltipraz, a number of ARE-associated enzymes were induced in a dose-dependent fashion [8]. In a separate study, 0.075% oltipraz in the diet reduced the volume fraction of AFB1-induced GGT+ foci at 3 months and completely prevented the occurrence of tumors for times up to two years [9]. Oltipraz given following AFB1 exposure also reduced the volume percent of GST-P+ foci [10].
It is not entirely clear whether oltipraz acts by increasing DNA repair. A cell-free system was used to examine nucleotide excision repair in human cells in vitro and in livers from rats exposed to AFB1 and oltipraz; NER DNA repair enzymes were not induced by oltipraz and are not known to be associated with AREs [11]. However, some studies suggest that eukaryotic NER expression is induced by exposure to phytochemicals (Gross-Steinmeyer et al. 2010), and alteration of DNA repair has been suggested as a pathway of chemoprevention for AFB1 carcinogenesis (Gross-Steinmeyer and Eaton, 2012). Oltipraz likely acts by activating the Nrf2-Keap1 pathway [12]. Oltipraz has been shown to be successful as a chemoprotectant in humans exposed to AFB1, presumably by similar mechanisms to those elucidated in rats [3,13,14].
Chlorophyllin
In Rainbow trout, chlorophyllin interferes with the formation of AFB1 adducts by forming a non-covalent complex with AFB1 [15]. This complex is not subject to metabolism by Phase 1 enzymes and thus the pro-mutagenic exo-epoxide metabolite cannot be formed. Chlorophyllin also induces ARE-associated enzymes HO-1 and NQO1 [16]; however, the primary mechanism for cancer prevention appears to be reduction in the uptake and bioavailability of AFB1 and its exo-epoxide [17,18]. In rats, administration of chlorophyllin after treatment with AFB1 had no effect on reducing the occurrence or volume of AHF in rats [19].
In human volunteers administered 14C-AFB1, co-administration of chlorophyllin increased the level of AFB1 equivalents in the urine [20]. This finding indicates that chlorophyllin works in humans in a similar fashion—by binding AFB1 prior to metabolism and thus limiting uptake/bioavailability.
Sulforaphane
Sulforaphane also appears to be a potent activator of the Nrf2-Keap1 pathway leading to increased expression of carcinogen detoxifying enzymes [12,21].
Other Phytochemicals
Green tea polyphenols and Korean red ginseng act as chemoprotectants. The effect is based on their anti-oxidant and anti-inflammatory properties, and likely mediated through the Nrf2-Keap1 anti-oxidant pathway, similar to what was described previously [22,23].
CDDO-Im and other Triterpenoids
Very recently, a landmark paper by Johnson et al. (2014) demonstrated that administration of the triterpenoid CDDO-Im prior to and during 28 days of dosing rats with AFB1 provided complete protection against the formation of both pre-neoplastic lesions and liver tumors. [24]. In addition, the rats treated with CDDO-Im maintained a reduced yet still significant burden of AFB1 adducts in their liver. Triterpenoids are potent inducers of the Nrf2-Keap1 ARE pathway. Commentaries on this paper noted both that Nrf2 activation introduced a nonlinearity into the AFB1 adduct-to-tumor relationship and also might provide significant protection in human populations exposed to AFB1 [2,25].
1. Yates MS, Kensler TW (2007) Keap1 eye on the target: chemoprevention of liver cancer. Acta Pharmacol Sin 28: 1331-1342.
2. Olden K, Vulimiri SV (2014) Laboratory to community: chemoprevention is the answer. Cancer Prev Res (Phila) 7: 648-652.
3. Kensler TW, Egner PA, Wang J-B, Zhu Y-R, Zhang B-C, et al (2004) Chemoprevention of hepatocellular carcinoma in aflatoxin endemic areas. Gastroenterology 127: S310-S318.
4. Ma Q, He X (2012) Molecular basis of electrophilic and oxidative defense: promises and perils of Nrf2. Pharmacol Rev 64: 1055-1081.
5. Jaramillo MC, Zhang DD (2013) The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev 27: 2179-2191.
6. Kwak MK, Itoh K, Yamamoto M, Sutter TR, Kensler TW (2001) Role of transcription factor Nrf2 in the induction of hepatic phase 2 and antioxidative enzymes in vivo by the cancer chemoprotective agent, 3H-1, 2-dimethiole-3-thione. Mol Med 7: 135-145.
7. Baldwin S, Parker RS (1987) Influence of dietary fat and selenium in initiation and promotion of aflatoxin B1-induced preneoplastic foci in rat liver. Carcinogenesis 8: 101-107.
8. Kensler TW, Egner PA, Dolan PM, Groopman JD, Roebuck BD (1987) Mechanism of protection against aflatoxin tumorigenicity in rats fed 5-(2-pyrazinyl)-4-methyl-1,2-dithiol-3-thione (oltipraz) and related 1,2-dithiol-3-thiones and 1,2-dithiol-3-ones. Cancer Res 47: 4271-4277.
9. Roebuck BD, Liu YL, Rogers AE, Groopman JD, Kensler TW (1991) Protection against aflatoxin B1-induced hepatocarcinogenesis in F344 rats by 5-(2-pyrazinyl)-4-methyl-1,2-dithiole-3-thione (oltipraz): predictive role for short-term molecular dosimetry. Cancer Res 51: 5501-5506.
10. Maxuitenko YY, MacMillan DL, Kensler TW, Roebuck BD (1993) Evaluation of the post-initiation effects of oltipraz on aflatoxin B1-induced preneoplastic foci in a rat model of hepatic tumorigenesis. Carcinogenesis 14: 2423-2425.
11. Sparfel L, Langouët S, Fautrel A, Salles B, Guillouzo A (2002) Investigations on the effects of oltipraz on the nucleotide excision repair in the liver. Biochem Pharmacol 63: 745-749.
12. Kensler TW, Roebuck BD, Wogan GN, Groopman JD (2011) Aflatoxin: a 50-year odyssey of mechanistic and translational toxicology. Toxicol Sci 120 Suppl 1: S28-S48.
13. Gross-Steinmeyer K, Eaton DL (2012) Dietary modulation of the biotransformation and genotoxicity of aflatoxin B(1). Toxicology 299: 69-79.
14. Sudakin DL (2003) Dietary aflatoxin exposure and chemoprevention of cancer: a clinical review. J Toxicol Clin Toxicol 41: 195-204.
15. Breinholt V, Schimerlik M, Dashwood R, Bailey G (1995) Mechanisms of chlorophyllin anticarcinogenesis against aflatoxin B1: complex formation with the carcinogen. Chem Res Toxicol 8: 506-514.
16. Zhang Y, Guan L, Wang X, Wen T, Xing J, Zhao J (2008) Protection of chlorophyllin against oxidative damage by inducing HO-1 and NQO1 expression mediated by PI3K/Akt and Nrf2. Free Radic Res 42: 362-371.
17. Hayashi T, Schimerlik M, Bailey G (1999) Mechanisms of chlorophyllin anticarcinogenesis: dose-responsive inhibition of aflatoxin uptake and biodistribution following oral co-administration in rainbow trout. Toxicol Appl Pharmacol 158: 132-140.
18. Breinholt V, Arbogast D, Loveland P, Pereira C, Dashwood R, et al (1999) Chlorophyllin chemoprevention in trout initiated by aflatoxin B(1) bath treatment: An evaluation of reduced bioavailability vs. target organ protective mechanisms. Toxicol Appl Pharmacol 158: 141-151.
19. Orner GA, Roebuck BD, Dashwood RH, Bailey GS (2006) Post-initiation chlorophyllin exposure does not modulate aflatoxin-induced foci in the liver and colon of rats. J Carcinog 5: 6. 20. Jubert C, Mata J, Bench G, Dashwood R, Pereira C, et al (2009) Effects of chlorophyll and chlorophyllin on low-dose aflatoxin B(1) pharmacokinetics in human volunteers. Cancer Prev Res (Phila) 2: 1015-1022. 21. Keum Y-S, Yu S, Chang PP-J, Yuan X, Kim J-H, et al (2006) Mechanism of action of sulforaphane: inhibition of p38 mitogen-activated protein kinase isoforms contributing to the induction of antioxidant response element-mediated heme oxygenase-1 in human hepatoma HepG2 cells. Cancer Res 66: 8804-8813.
22. Qin G, Ning Y, Lotlikar PD (2000) Chemoprevention of aflatoxin B1-initiated and carbon tetrachloride-promoted hepatocarcinogenesis in the rat by green tea. Nutr Cancer 38: 215-222.
23. Kim Y-S, Kim Y-H, Noh J-R, Cho E-S, Park J-H, Son H-Y (2011) Protective Effect of Korean Red Ginseng against Aflatoxin B1-Induced Hepatotoxicity in Rat. J Ginseng Res 35: 243-249.
24. Johnson NM, Egner PA, Baxter VK, Sporn MB, Wible RS, et al (2014) Complete protection against aflatoxin B1-induced liver cancer with triterpenoid: DNA adduct dosimetry, molecular signature and genotoxicity threshold. Cancer Prev Res (Phila) .
25. Eaton DL, Schaupp CM (2014) Of mice, rats, and men: could Nrf2 activation protect against aflatoxin heptocarcinogenesis in humans? Cancer Prev Res (Phila) 7: 653-657.
Evidence Assessment
Support for Biological Plausibility of KERs
| Support for Biological Plausibility of KERs | Defining Question | High (Strong) | Moderate | Weak |
| a) Is there a mechanistic (i.e., structural or functional) relationship between KEup and KEdown consistent with established biological knowledge? | Extensive understanding of the KER based on extensive previous documentation and broad acceptance (e.g., mutation leading to tumors)
-Established mechanistic basis |
The KER is plausible but scientific understanding is not completely established. | Only limited or indirect evidence for KER (i.e., based on empirical support, only (See 3.) | |
| Hepatic metabolic activation directly to formation of pro-mutagenic DNA adducts | Biological Plausibility of the pre-MIE => MIE is Strong
Rationale: Long-established knowledge of the metabolism of AFB1 to specific reactive electrophiles that form pro-mutagenic DNA adducts. |
|||
| Pro-mutagenic adduct formation directly to Insufficient / Mis-repair of pro-mutagenic adducts | Biological Plausibility of MIE => KE1 is Strong.
Rationale: not much direct empirical support but strongly accepted. |
|||
| Insufficient/Mis-repair directly to Induced mutations in critical gene | Biological Plausibility of KE1 => KE2 is Strong
Rationale: Long established knowledge : Empirical data from yeast with defective repair systems leads to increased mutations—infer increased mutations in critical genes |
|||
| Induced mutations directly to clonal expansion / cell proliferation (AHF) | Biological Plausibility of KE2 => KE3 is Strong
Rationale: Necessary. Based on chemoprevention studies, HBV, and the plethora of initiation-promotion studies. |
|||
| Clonal Expansion / Cell Proliferation directly to HCC | Biological Plausibility of KE3 => AO is Strong
Rationale: Long established knowledge; the plethora of longer term initiation-promotion studies provide much evidence of the link from clonal expansion of foci to HCC. |
|||
| Indirect: Pro-mutagenic adducts to clonal expansion / cell proliferation (AHF) | Biological Plausibility of MIE => KE3 is Strong
Rationale: Based on the relationship of adducts to AHF, data on chemopreventive agents that specifically decrease adduct formation also decrease the occurrence of AHF. |
|||
| Indirect: Pro-mutagenic adducts to HCC | Biological Plausibility of MIE => AO is Strong
Rationale: The relationship of adducts to HCC depends of two well-established relationships between adducts and AHF and between AHF and HCC. Because of these well-estabilished relationships, the biological plausibility is judged to be strong. |
|||
| Indirect: Mutations to HCC | Biological Plausibility of KE2 => AO is Strong
Rationale: The relationship of mutations to cancer is well-established. However, what is not clear is whether mutations observed early in the cancer process are the same as those observed in tumors. However, the relationship of adducts to AHF and AHF to tumors are both strong . thusthis indirect KER is also strong. |
|||
Dose-Time Concordance Table Based on Available Data
AFB1 is a data-rich chemical, and it is clear that it induces hepatocellular carcinoma (HCC) in rats, fish, and in humans, among other species. The vast majority of experiments, however, were not designed with the intention of arraying the information in a dose- and time-concordant manner. As indicated in this table, there are no quantitative data (aside from the induction of pre-neoplastic lesions and HCCs) that can be used to develop the quantitative relationships between the KEs. The KEs that have been identified for AFB1-induced HCC begin with metabolism to reactive moieties and a key metabolite forming pro-mutagenic DNA adducts. If these adducts are not repaired, are mis-repaired, or are insufficiently repaired, this can lead to the induction of mutations (primarily G:C to T:A transversions) across the genome (Dycaico et al., 1996; Chen et al. 2010 ). There are sufficient data to indicate that at least one hot-spot for mutation induction occurs in codon 249 (G:C to T:A transversion) of the p53 gene. This conclusion is based on the fact that this mutation has been observed in up to 75% of liver cancers in humans residing in the high AFB1 incidence areas of China and East Africa, but the same mutation is rarely seen in no or low AFB1 areas (Hsu et al., 1991 and Bressac et al., 1991). Because of a lack of practicable methods, there are no studies conclusively demonstrating that AFB1 exposure induces these specific mutations in vivo. To be conclusive, demonstration of specific cancer gene mutation would have to be shown following relatively short exposure to AFB1 and long before the observation of any pre-neoplastic AHF lesions or HCCs.
There have, however, been some studies in which rodents have been exposed to AFB1 and either pro-mutagenic DNA adducts or increased mutant frequency in a surrogate gene (Lac I or cII) of a transgenic rodent have been observed. These data are shown in the table. There is some dose-response information for the AFB1 DNA adduct formation, but the rodent studies were conducted with only a single mutagenic dose. It should be noted that the neonatal mouse is very sensitive to the induction of mutations (and also susceptible to the induction of tumors), but the adult mouse exposed to AFB1 shows neither mutation nor tumors; this difference in sensitivity is believed to be due to differential ontological expression patterns in glutathione-S-transferase (GST) activity.
Data from studies with chemoprevention agents add considerably to the WOE and understanding of the KE as necessary but not sufficient. In particular, two datasets are shown in the Data Concordance Table: Roebuck et al. (1991) with oltipraz, and Johnson et al. (2014) with CDDO-Im. Both datasets demonstrated a significant reduction in tumor burden with a concomitant decrease in altered hepatic foci (AHF) (Johnson et al., 2014), that was correlated with reduction but not elimination of the pro-mutagenic DNA adducts. The reduction in DNA adducts was likely due in part to induction of GST activity. In any case, both datasets demonstrate that the presence of pro-mutagenic adducts, alone, does not result in AHF or HCC; that is, adduct formation is necessary but not sufficient.
| Dose-Time Concordance Table Based on Available Data | Increasing Time --> | ||||||
| Dose (ppb in diet) | Met. Activ'n | Pro-mutagenic DNA Adducts | Insufficient/Mis-repair of pro-mutagenic DNA adducts* | Induced mutation in critical gene(s)** | Clonal expansion of mutant cells (pre-neoplastic lesions) | Hepatocellular Carcinomas | |
| 0 | -/+ (0.06)* | --- (0) | |||||
| 1 | + (0.32) | -/+ (0.09) | |||||
| 5 | + (0.23) | -/+ (0.05) | |||||
| 15 | ++(0.62) | +(0.19) | |||||
| 50 | ++ (0.60) | +++ (0.8) | |||||
| 100 | ++(0.43) | ++++(1.0) | |||||
Data from same study (Wogan et al., 1974, Table 1)
- Wogan et al., 1974, Table 1. [Wogan G.N., Paglialunga S., Newberne P.M. 1974. Carcinogenic effects of low dietary levels of aflatoxin B1 in rats, Food. Cosmet. Toxicol. 12, 681-68].
| Increasing Time --> | |||||||
| Dose (Rat Liver mg/kg ip, Single Dose) | Met. Activ'n | Pro-mutagenic DNA Adducts | Insufficient/Mis-repair of pro-mutagenic DNA adducts* | Induced mutation in critical gene(s)** | Clonal expansion of mutant cells (pre-neoplastic lesions) | Hepatocellular Carcinomas | |
| 10 | 0.37 pmol adduct/mg DNA | ||||||
| 25 | 0.48 pmol adduct/mg DNA | ||||||
| 65 | 1.47 pmol adduct/mg DNA | ||||||
| 160 | 3.93 pmol adduct/mg DNA | ||||||
| 390 | 8.54 pmol adduct/mg DNA | ||||||
| 1000 | 16.48 pmol adduct/mg DNA | ||||||
| Laci/Big Blue Rat*** mg/kg/d ip | |||||||
| 0.25 | 500 mutants/ 106 (measured in a surrogate gene) | ||||||
| cII/Big Blue mouse ip**** | |||||||
| 6 mg/kg (neonate) | 900 mutants/ 106 (measured in a surrogate gene) | ||||||
| 6 mg/kg (adult) | no increase | ||||||
| 60 mg/kg (adult) | no increase | ||||||
Table of Reversibility of KEs
| Dose-Time Concordance Table for Reversibility of KEs | Increasing Time --> | ||||||
| Dose (Rat Liver mg/kg ip, Single Dose) | Met. Activ'n | Pro-mutagenic DNA Adducts | Insufficient/Mis-repair of pro-mutagenic DNA adducts* | Induced mutation in critical gene(s)** | Clonal expansion of mutant cells (pre-neoplastic lesions) | Hepatocellular Carcinomas | |
| AFB1 w/o Oltipraz | 0.25 mg/kg | +++ | -/+ 13% | -/+ 11% | |||
| AFB1 w/ Oltipraz | 0.25 mg AFB1/kg w/ 0.075% in diet | + 75% reduction | -/+ 4% | -/+ 0% | |||
| AFB1 w/o CDDO-Im^^ | 200 mg/kg | +++ | +++++ (23/23) | ++++ (96%, 22/23) | |||
| AFB1 w/ CDDO-Im^^ | 200 mg/kg | + (N7: 50-80% reduction FAPy: 50-70% reduction) | +(3/20) | --(0/20) | |||
Table References Chen T., Heflich R.H., Moore M.M., Mei N.2010. Differential mutagenicity of aflatoxin B1 in the liver of neonatal and adult mice.. Environ Mol Mutagen. 51(2), 156-63
Dycaico M.J., Stuart G.R., Tobal G.M., de Boer J.G., Glickman B.W., Provost G.S.1996. Species-specific differences in hepatic mutant frequency and mutational spectrum among lambda/lacI transgenic rats and mice following exposure to aflatoxin B1.. Carcinogenesis. 17(11), 2347-5
Johnson N.M., Egner P.A., Baxter V.K., Sporn M.B., Wible R.S., Sutter T.R., Groopman J.D., Kensler T.W., Roebuck B.D.2014. Complete protection against aflatoxin B1-induced liver cancer with triterpenoid: DNA adduct dosimetry, molecular signature and genotoxicity threshold.. Cancer Prev Res (Phila).
Roebuck B.D., Liu Y.L., Rogers A.E., Groopman J.D., Kensler T.W.1991. Protection against aflatoxin B1-induced hepatocarcinogenesis in F344 rats by 5-(2-pyrazinyl)-4-methyl-1,2-dithiole-3-thione (oltipraz): predictive role for short-term molecular dosimetry.. Cancer Res. 51(20), 5501-6
Hypothetical and Ideal Data: Dose-Time Concordance Table
Given the importance of evaluating AOP information using a dose- and time-concordance table, and the fact that the appropriate data are not available for AFB1, we constructed a table to indicate what would be expected in a hypothetical ideal situation. It is expected that each of the KE will show a dose-response relationship with upstream KE observed at lower exposure levels than the downstream KE, and the AO observed only at the higher doses. Likewise the upstream KE would be expected to be observed at earlier times than the downstream KE or the apical outcome. Thus metabolism to reactive AFB1 forms must occur prior to the observation of DNA adducts. These adducts must be insufficiently repaired, not repaired, or mis-repaired prior to the observation of mutation in the critical cancer gene. For a mutagenic AOP, it is important that the mutation is induced prior to the clonal expansion of cells. Furthermore in this AOP a critical gene mutation leads to the formation of pre-neoplastic lesions, which must precede the formation of tumors. For chronic exposure, one would expect that the frequency of mutant cells would increase with the increased length of exposure; that is, the early KE will continue to occur with time leading to more adducts being formed and more mutations being induced.
While the studies presented in the concordance did not specifically measure DNA repair or induction of mutations, they do provide data on KE#3 (AHF frequency) and the AO (HCC tumor incidence). Treatment with chemoprotective agents decreased AHF formation significantly for oltipraz (Roebuck et al., 1991), while for CDDO-Im, AHF incidence was reduced to background/control levels (Johnson et al., 2014). In these studies, the AO HCC tumor incidence was either significantly reduced (oltipraz) or completely eliminated (CDDO-Im; 0% HCC incidence).
| Dose | Increasing Time --> | ||||||
| KE (ppb in diet) | Metabolic Activation | Pro-mutagenic DNA Adducts | Insufficient/Mis-repair of pro-mutagenic DNA adducts | Induced mutation in critical gene(s) | Clonal expansion of mutant cells (pre-neoplastic lesions) | Hepatocellular Carcinomas | |
| 0 | - | - | - | - | -/+ (0.06) | --- (0) | |
| 1 | + | ++ | + | + | + (0.32) | -/+ (0.09) | |
| 5 | ++ | +++ | ++ | ++ | + (0.23) | -/+ (0.05) | |
| 15 | +++ | ++++ | +++ | +++ | ++ (0.62) | + (0.19) | |
| 50 | ++++ | +++++ | +++++ | +++++ | ++ (0.60) | +++ (0.8) | |
| 100 | +++++ | ++++++ | +++++ | +++++ | ++ (0.43) | ++++ (1.0) | |
| Reversibility: | |||||||
| **AFB w/o Oltipraz | +++ | +++ | ++ | ++ | -/+ | -/+ | |
| **AFB w/ Oltipraz | +++ | + | + | + | -/+ | -/+ | |
| ***AFB w/o CDDO-Im | +++ | +++++ | ++++ | ||||
| ***AFB w/ CDDO-Im | + | + | -- | ||||
* Wogan et al., 1974, Table 1. Wogan G.N., Paglialunga S., Newberne P.M. 1974. Carcinogenic effects of low dietary levels of aflatoxin B1 in rats, Food. Cosmet. Toxicol. 12, 681-68.
|
|||||||
Known Modulating Factors
Quantitative Understanding
Support for Biological Plausibility of KERs
| Empirical Support for KERs | Defining Question | High (Strong) | Moderate | Weak |
| Does the empirical evidence support that a change in KEup leads to an appropriate change in KEdown? Consider dose-response concordance, temporality (i.e., KEup occurs at lower dose and earlier time point than KE down and incidence concordance (i.e., is the incidence of KEup > than that for KEdown). | Multiple studies showing change in both events following exposure to a specific stressor. (Extensive evidence for temporal, dose-response and incidence concordance. | There is at least one line of experimental evidence showing change in both events following exposure to a specific stressor.There is at least one line of experimental evidence for temporal and/or dose-response and incidence concordance. | Limited or no studies reporting change in both events following exposure to a specific stressor (i.e., endpoints never measured in the same study or not at all).
-Lacking evidence of temporal or dose-response concordance. |
|
| Hepatic metabolic activation directly to formation of pro-mutagenic DNA adducts | Empirical Support of the pre-MIE => MIE is Strong
Rationale: Pre-MIE (upstream) occurs prior to MIE (downstream) so temporal concordance strong; without metabolic activation, do not get mutations with in vitro systems—no activation, no adducts to lead to mutations |
|||
| Pro-mutagenic adduct formation directly to Insufficient / Mis-repair of pro-mutagenic adducts | Empirical Support of the MIE => KE1 is Moderate
Rationale: Data are indirect, such as yeast with DNA repair deficiencies demonstrate increased mutations (Guo et al., 2005), but theory is well accepted; also indirectly supported by data demonstrating decreases in pro-mutagenic adducts result in decreased downstream events (AHF & HCC) (Roebuck et al., 1991; Johnson et al., 2014). |
|||
| Insufficient/Mis-repair directly to Induced mutation in critical genes | Empirical Support of the KE1 => KE2 is Moderate
Rationale: While there is no available information that insufficient/mis-repair of DNA adducts leads directly to induced mutations in critical genes. there is evidence that AFB1 induces mutations in a number of different mutation models, both in vitro and in vivo. The induction of mutations occurs in a dose-response manner and also occurs shortly after the exposure to AFB1 Based on data from the surrogate gene mutation assays, it is clear that AFB1 is able to induce the type of mutation that is seen in codon 249 of the p53 gene in human HCCs from regions with high AFB1 exposure |
|||
| Induced mutations directly to clonal expansion/proliferation of mutant cells (AHF) | Empirical Support of the KE2 => KE3 is Moderate
Rationale: Assume critical gene mutation is extant resulting in transformed cells, typically identified with altered enzymatic/histopathologic techniques; KER indirectly supported by <1% GST-P+ to >10% GST-P+ hepatocytes after 3-4 wks of AFB1 as surrogate for mutation in AHF mutant cells, as stated in Johnson et al., 2014; Plausibility based on other models of HCC by mutagenic: induction and progression of AHF. |
|||
| Clonal Expansion directly to HCC | Empirical Support of the KE3 => AO is Strong: pathogenesis of AHF to liver cancer (Dragan & Pitot)
Rationale: The plethora of longer term initiation-promotion studies provide much evidence of the link from clonal expansion of foci to HCC. Indirect for AFB but may have plenty of evidence |
|||
| Indirect: Pro-mutagenic adduct to AHF | Empirical Support of MIE => KE3 is Strong: chemoprevention data with reduced adducts & decrease/no foci
Rationale Based on the relationship of adducts to AHF, data on chemopreventive agents that specifically decrease adduct formation also decrease or even eliminate the occurrence of AHF. |
|||
| Indirect: Pro-mutagenic adduct to HCC | Empirical Support of MIE => AO is Strong: chemoprevention data with reduced adducts & decrease/no HCC
Rationale: AFB1 appears to act as both a tumor initiator and a tumor promoter. The tumor promotional effects are blocked by triterpenoid chemopreventants and decrease tumor occurrence in the presence of adducts. Hence, adducts are necessary for tumors to occur but not entirely sufficient. |
|||
| Indirect: Mutations to HCC | Empirical Support of KE2 => AO is Moderate:
Rationale: Strong evidence that AFB1 preferentially induces G:C to T:A transversions in a variety of mutation assays; Codon 249 3rd base G:C to T:A transversions found in up to 50-75% human tumors from populations with high AFB1 exposure & low HBV incidence; G:C to T:A transversion mutations matches AFB1-induced mutations; no info on whether those tumor mutations were early initiating mutations or late-forming, progression-related mutations, so evidence is moderate; biological plausibility is strong |
|||
Uncertainties and Conflicting Evidence for KERs and AOP
Support for Biological Plausibility of KERs
| Uncertainty and Conflicting Evidence for KERs and AOP | Defining Question | High (Strong) | Moderate | Weak |
| Are there inconsistencies in empirical support across taxa, species which don’t align with appropriate pattern for hypothesized KERs and AOP?
Are there significant knowledge gaps or uncertainties with regard to the relationship between the KEs and overall AOP? |
No (or very few) knowledge gaps or inconsistent / conflicting lines of evidence.) | Some inconsistent evidence but which can be explained by factors such as experimental design, technical considerations, differences among laboratories, etc.) | Contradictory evidence in for which no plausible explanation is known. | |
| Hepatic metabolic activation directly to formation of pro-mutagenic DNA adducts | Inconsistencies / Uncertainties of Pre-MIE => MIE is Strong
Rationale: Highly certain. |
|||
| Pro-mutagenic adduct formation directly to Insufficient / Mis-repair of pro-mutagenic adducts | Inconsistencies / Uncertainties of MIE => KE1 is Strong.
Rationale: Highly certain. |
|||
| Insufficient/Mis-repair directly to Induced mutations in critical gene | Inconsistencies / Uncertainties of KE1 => KE2 is Moderate
Rationale: No direct evidence/data on induction of mutation in critical gene; AFB1 induces G:C to A:T transversion mutations in variety of mutation assays (surrogate genes); high percentage of tumors in high AFB1 exposure regions have matching G:C to A:T transversion mutations |
|||
| Induced mutations directly to clonal expansion of mutant cells | Inconsistencies / Uncertainties of KE2 => KE3 is Moderate.
Assume critical gene mutation is extant resulting in transformed cells, typically identified with altered enzymactic/histopath techniques; KER possibly supported by <1% GST-P+ to >10% GST-P+ hepatocytes after 3-4 wks of AFB1 as surrogate for mutation in AHF mutant cells, as stated in Johnson et al., 2014; Plausibility based on other models of HCC by mutagenic: induction and progression of AHF Rationale: Plausible but with minimal knowledge support due to technical limitations |
|||
| Clonal Expansion directly to HCC | Inconsistencies / Uncertainties of KE3 => AO is Strong
Rationale: Chemoprevention data: no foci leads to no tumors; Dragan & Pitot liver tumor etiology |
|||
| Indirect: Pro-mutagenic adduct to AHF | Inconsistencies / Uncertainties of MIE => KE3 is Strong
Rationale: It is not known whether the early occurring mutations that occur from N7 or FAPy adducts are the same as those that lead to AHF. Nonetheless, the effects of mutations are apparent in AHF and used for their identification. |
|||
| Indirect: Pro-mutagenic adduct to HCC | Inconsistencies / Uncertainties of MIE => AO is Strong
Rationale: It is not known whether the adducts occurring from AFB1 exposure lead to mutations that are directly causal for tumors. Nonetheless, there is a strong relationship between adducts and tumors when chemopreventants are not used; conversely decreases in adduct levels reduced or eliminated HCC incidence. |
|||
| Indirect: Critical Gene Mutations to HCC | Inconsistencies / Uncertainties of KE2 => AO is Moderate
Rationale: There are no data to directly address this. Indirect evidence: There is a high association between exposure to AFB1 and codon 249 mutations in human HCC. |
|||
Considerations for Potential Applications of the AOP (optional)
References
Jarabek A.M., Pottenger L.H., Andrews L.S., Casciano D., Embry M.R., Kim J.H., Preston R.J., Reddy M.V., Schoeny R., Shuker D., Skare J., Swenberg J., Williams G.M., Zeiger E.2009. Creating context for the use of DNA adduct data in cancer risk assessment: I. Data organization.. Crit Rev Toxicol. 39(8), 659-7
Moore, M.M., R.H. Heflich, L. Haber, B.C. Allen, A. M. Shipp, and R.L. Kodell. 2008. Analysis of in vivo mutation data can inform cancer risk assessment, Regulatory Toxicology and Pharmacology. 51:151-161.
Parsons BL, Myers MB, Meng F, Wang Y, McKinzie PB. 2010. Oncomutations as biomarkers of cancer risk. Environ Mol Mutagen. 51(8-9):836-850.
Pottenger L.H., Andrews L.S., Bachman A.N., Boogaard P.J., Cadet J., Embry M.R., Farmer P.B., Himmelstein M.W., Jarabek A.M., Martin E.A., Mauthe R.J., Persaud R., Preston R.J., Schoeny R., Skare J., Swenberg J.A., Williams G.M., Zeiger E., Zhang F., Kim J.H., et al.2014. An organizational approach for the assessment of DNA adduct data in risk assessment: case studies for aflatoxin B1, tamoxifen and vinyl chloride.. Crit Rev Toxicol. 44(4), 348-91