
This AOP is licensed under the BY-SA license. This license allows reusers to distribute, remix, adapt, and build upon the material in any medium or format, so long as attribution is given to the creator. The license allows for commercial use. If you remix, adapt, or build upon the material, you must license the modified material under identical terms.
AOP: 15
Title
Alkylation of DNA in male pre-meiotic germ cells leading to heritable mutations
Short name
Graphical Representation
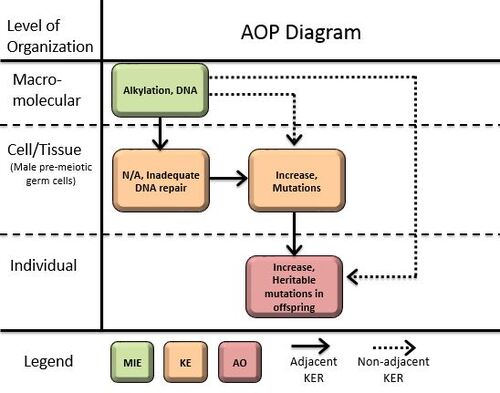
Point of Contact
Contributors
- Carole Yauk
Coaches
OECD Information Table
| OECD Project # | OECD Status | Reviewer's Reports | Journal-format Article | OECD iLibrary Published Version |
|---|---|---|---|---|
| 1.11 | WPHA/WNT Endorsed |
This AOP was last modified on April 29, 2023 16:02
Revision dates for related pages
| Page | Revision Date/Time |
|---|---|
| Increase, Heritable mutations in offspring | November 29, 2016 19:06 |
| Alkylation, DNA | September 16, 2017 10:14 |
| Increase, Mutations | May 15, 2023 08:47 |
| Inadequate DNA repair | March 08, 2024 12:15 |
| Alkylation, DNA leads to Inadequate DNA repair | December 10, 2019 10:43 |
| Inadequate DNA repair leads to Increase, Mutations | March 08, 2024 15:00 |
| Alkylation, DNA leads to Increase, Mutations | November 29, 2016 19:53 |
| Alkylation, DNA leads to Increase, Heritable mutations in offspring | November 29, 2016 19:53 |
| Increase, Mutations leads to Increase, Heritable mutations in offspring | November 29, 2016 19:59 |
Abstract
Germ cell/heritable mutations are important regulatory endpoints for international agencies interested in protecting the health of future generations. However, germ cell mutation analysis has been hampered by a lack of efficient tools. With the publication of the OECD test guideline TG488 (rodent transgene mutation assay) and new technologies (including next generation sequencing) this field is experiencing renewed focus. Indeed, regulatory approaches to assess germ cell mutagenicity were the focus of an IWGT workshop (Yauk et al., 2013). Of particular concern is the inability to address this endpoint through high-throughput screening assays (because spermatogenesis cannot be carried out in culture), and mutagenesis is an important gap in existing high-throughput tests. The motivation for developing this AOP was to provide context for new assays in this field, identify research gaps and facilitate the development of new methods.
In this AOP, a compound capable of alkylating DNA is delivered to the testes causing germ cell mutations and subsequent mutations in the offspring of the exposed parents. The AOP requires uptake of the parent compound or metabolite in spermatogonia and interaction with DNA in those cells. DNA alkylation in male pre-meiotic germ cells is the molecular initiating event. A variety of different DNA adducts are formed that are subject to DNA repair; however, at high doses the repair machinery becomes saturated or overwhelmed. The fate of remaining adducts includes: (1) attempted DNA repair by alternative DNA repair machinery, or (2) no repair. Key event (KE) 1 is insufficient or incorrect DNA repair. Lack of repair can lead to replication of adducted DNA and ensuing mutations in male pre-meiotic germ cells (KE2). Mutations that do not impair spermatogenic processes will persist in these cells and eventually be present in the mature sperm. Thus, the mutations can be transmitted to the offspring (adverse outcome – inherited mutations). It is well documented that mice and other animals exposed to alkylating agents develop mutations in male pre-meiotic germ cells that are then found in sperm, resulting in the transmission of mutations to their offspring. There is a significant amount of empirical evidence supporting the AOP and the overall weight of evidence is strong. Although there are some gaps surrounding some mechanistic aspects of this AOP, the overarching AOP is widely accepted and applies broadly to any species that produces sperm.
AOP Development Strategy
Context
De novo germ cell mutations are changes in the DNA sequence of sperm or egg that can be inherited by offspring. De novo mutations contribute to a wide range of human disorders including cancer, infertility, autism, schizophrenia, intellectual disability, and epilepsy (Girirajan et al. 2010; Hoischen et al. 2010; Ku et al. 2012; Lupski 2010; Morrow 2010; Vissers et al. 2010). Each child inherits, on average, approximately one de novo mutation per 100 million nucleotides delivered via the parental egg and sperm (Conrad et al. 2011; Kong et al. 2012; O'Roak et al. 2012; Roach et al. 2010). The precise locations and types of mutations in the genomic DNA sequence govern the outcome of these mutations (e.g., protein coding versus intergenic sequences, conserved versus non-conserved mutations, etc.). Although a large portion of human DNA is of unknown function, recent literature suggests that at least 80% of the genome is transcribed, and most DNA is expected to have a biological function (Bernstein et al. 2012). It has been estimated that the proportion of coding and splice-site base substitutions that result in truncating mutations is ~5% (Kryukov et al. 2007), and that as many as 30% of missense mutations are also likely to be highly deleterious due to loss of function (Boyko et al. 2008). When they occur in functional sites, de novo mutations can cause embryonic or fetal lethality, or if viable, can produce a broad spectrum of inherited genetic disorders. Recent estimates suggest that a human genome contains approximately 100 loss-of-function variants, with as many as 20 exhibiting complete loss of gene function (McLaughlin et al. 2010). Therefore, de novo mutations contribute to the overall population genetic disease burden. The present AOP focuses on DNA alkylation in spermatogonia that causes inherited mutation transmitted via sperm, arguably one of the most well characterized modes of action in genetic toxicology. Humans are exposed to alkylating agents from external (e.g., abiotic plant materials, tobacco smoke, combustion products, chemotherapeutic agents) and internal (e.g., byproducts of oxidative damage and cellular methyl donors) sources.
Strategy
Summary of the AOP
Events:
Molecular Initiating Events (MIE)
Key Events (KE)
Adverse Outcomes (AO)
| Type | Event ID | Title | Short name |
|---|
| MIE | 97 | Alkylation, DNA | Alkylation, DNA |
| KE | 155 | Inadequate DNA repair | Inadequate DNA repair |
| KE | 185 | Increase, Mutations | Increase, Mutations |
| AO | 336 | Increase, Heritable mutations in offspring | Increase, Heritable mutations in offspring |
Relationships Between Two Key Events (Including MIEs and AOs)
| Title | Adjacency | Evidence | Quantitative Understanding |
|---|
| Alkylation, DNA leads to Inadequate DNA repair | adjacent | High | Moderate |
| Inadequate DNA repair leads to Increase, Mutations | adjacent | High | Moderate |
| Increase, Mutations leads to Increase, Heritable mutations in offspring | adjacent | High | Moderate |
| Alkylation, DNA leads to Increase, Mutations | non-adjacent | High | Moderate |
| Alkylation, DNA leads to Increase, Heritable mutations in offspring | non-adjacent | High | Moderate |
Network View
Prototypical Stressors
Life Stage Applicability
| Life stage | Evidence |
|---|---|
| Adult | High |
Taxonomic Applicability
Sex Applicability
| Sex | Evidence |
|---|---|
| Male | High |
Overall Assessment of the AOP

Before developing this AOP a review of the literature was undertaken to identify studies in which male germ cells were exposed to alkylating agents and measures of DNA adducts, DNA repair and mutations, as well as mutations in offspring, were evaluated. The focus of this AOP (as described in the KERs) is on O-alkylating agents, which are signficantly more mutagenic than N-alkylation chemicals. Studies where sufficient information relating to the chemicals used, dose, tissue, time-point, animal model, experimental procedures and experimental results were available were considered to assess empirical data in germ cells for each of the KEs and KERs in the AOP. The germ cell database on which the AOP was based is found in Supplementary Table I (SupplementalTablesFigures) and is comprised of 32 studies. No study measured multiple KEs within it; however, for each KE there were at least two dose-response and time-series analyses for at least one alkylating agent. We consider this overall number of high quality studies to be fairly extensive evidence of the ability of O-alkylating agents to cause adducts and mutations in germ cells, and mutations in offspring, although no studies were ideally suited to establish the empirical linkages between the KERs. We thus compared results across studies where possible to attempt to do this. All of the studies either used ENU as the primary study compound, or applied ENU as one of the positive controls to assess other alkylating agents. Strong dose-response data for mutations occurring in exposed pre-meiotic germ cells and mutations in offspring are only available for ENU. The other alkylating agents show varying degrees mutagenicity, but single doses were used in most studies. Thus, the evaluation of concordance of the dose-response could only be undertaken with ENU for in vivo germ cell and heritable effects. However, where possible we used information from research on somatic cells to provide additional support for the KERs. In particular, experiments in somatic cells were necessary to assess the involvement of DNA repair in removing adducts and preventing mutations. Overall, we note that the rationale for claiming high confidence in this AOP and its KERs is based primarily on the more influential Bradford Hill consideration of biological plausibility, with decades of research having been done in somatic and germ cells on DNA damage, repair and mutation. Much of the data, then, supporting AOP evaluation derives from historical studies from the 1990’s, with less recent evidence. As noted, a primary motivation for developing this AOP was the recent release of TG 488, and newly available whole generation sequencing methods, which we expect to be increasingly applied. Thus, additional well-designed experiments that dissect the relationships between alkyl adducts, mutations in sperm, and mutations in offspring to assess essentiality and empirical support are expected in the future through application of these improved approaches. Below we describe each KE and KER in detail, using the wiki entries as a guide to the order of presentation and the content described.
Domain of Applicability
This AOP is relevant exclusively to mature males and their pre-meiotic germ cells. Although not considered in this AOP, progenitor germ cells from earlier life stages may also be susceptible to induced mutations from alkylating agents, which could then be transmitted to offspring after sexual maturity. Relevant endpoints have been characterized across different taxa: (1) alkyl adduct levels in this AOP were from hamsters, mice and rats; (2) repair of alkylated DNA has been studied in prokaryotes to higher eukaryotes, including human cells in culture (while there are differences across taxa, all species have some DNA repair systems in place and it is common to extrapolate conclusions across eukaryotic species); (3) mutations in male germ cells were measured in mice and fish; and (4) mutations in offspring were measured in Drosophila, Japanese Medaka and mice. Quite generally, the AOP applies to any species that produces sperm. The similarity in spermatogenesis and in DNA repair of alkyl adducts is well documented across rodents and humans (Adler 1996). Heritable mutations are the basis of evolution and occur in every species. That mutations in sperm are transmitted to offspring in humans is best demonstrated by studies exploring the effects of ageing. Significant increases are observed in the amount of DNA damage and mutation as human males age (reviewed in Paul and Robaire 2013). Similarly, increased incidence of single nucleotide mutations and microsatellite mutation in the offspring of ageing fathers has recently been measured by advanced genomics technologies (Kong et al. 2012; Sun et al. 2012). Lifestyle factors including smoking and lower income brackets in human fathers in associated with increased minisatellite mutations in their offspring (LinSchooten et al., 2013).
Essentiality of the Key Events
Essentiality was not directly tested for all of the KEs. The MIE cannot be ‘blocked’ in any way to our knowledge (e.g., as you might block a receptor-binding MIE). However, as described in the KERs, enhanced DNA repair of alkylated DNA reduces mutation frequencies and reduction in repair increases mutation frequencies, supporting the essentiality of KE1 (i.e., moderate support). Correct repair of the alkylated DNA (i.e., a block of KE1) will not lead to mutation. For example, MGMT overexpression protects mgt1 mutant yeast against alkylation-induced mutation (Xiao and Fontanie 1995). In addition, Big Blue® mice over-expressing human AGT exhibit greatly reduced O6-methylguanine-mediated lacI and K-ras mutations in the thymus following treatment with MNU (Allay et al. 1999) relative to wild type Big Blue® mice. Insufficient DNA repair is well-established to lead to mutations. In addition, inactivation of MGMT sensitizes cells to alkylation-induced mutagenesis resulting in an increased number of mutations per adduct (Thomas et al. 2013).
The remainder of the AOP requires transmission of mutations in sperm to offspring. There are no means to study the essentiality of mutations in sperm. Once mutations occur in male pre-meiotic germ cells, they cannot be removed to observe whether occurrence in offspring is decreased. In addition, mutations that occur in stem cells are propagated clonally and can become fixed in the spermatogonial cell population. Thus, waiting a longer period of time, or removing the exposure, is not effective in causing a decline in the mutation frequency. Therefore, the essentiality of this KE is inferred by the biology of the pathway and cannot be addressed directly with experimental evidence.
Evidence Assessment
Biological plausibility of the KERs: Strong. There is extensive understanding of the ability of alkylating agents to cause DNA adducts, the requirement for overcoming DNA repair, and the resulting mutations that arise in both somatic and germ cells. It is established that exposure to alkylating agents produced mutations in germ cells – ENU is used in genetic screening to produce mutations to derive new phenotypes for research.
Empirical support for the KERs: Across the KERs the degree of support ranges from weak to strong (File:AssessmentSummaryAop-15.pdf - Table II). Support from somatic cells in culture contributes to moderate calls for the relationships between adduct formation, insufficient DNA repair and mutation. The weak call is based on lack of empirical data to support that mutations in germ cells are transmitted to offspring. However, increased mutation frequencies in germ cells occur following exposure to the same types of chemicals that cause increased mutations in the offspring. It should be noted that biological plausibility for this KER is strong as it is based on understanding of molecular biology and evolution. The strongest support is associated with the indirect KER linking alkylation of DNA to mutation in germ cells (KER4). This is primarily based on extensive evidence in both somatic and germ cells demonstrating that chemicals that alkylate DNA cause mutations, that alkyl adducts occur at a greater incidence than mutations at matching doses, and that alkyl adducts precede mutations. In somatic cells, work has been done on many different chemicals, whereas the germ cell data were primarily for the chemical ENU (but data were also available for a few select other chemicals) (File:AssessmentSummaryAop-15.pdf - Table I, Figure 2). In addition, data are available for multiple species to support this indirect KER. There is a large degree of consistency in the germ cell literature to show that a variety of O-alkylating agents cause male germ cell mutations in many species (Drosophila, fish and rodent) and that these effects occur at many mutational loci (e.g., mutations in genes that are inherited measured with the Specific Locus Test, sperm mutations in tandem repeat DNA sequences, tandem repeat mutations in offspring, transgene mutations in sperm). Many alkylating agents have been tested to show that they create adducts in male rodent germ cells (e.g., DEN, ENU, EMS, DES), mutations in male mouse germ cells (ENU, IPMS and MNU) and mutations in the offspring of exposed male mice (ENU, MNU and IPMS). In summary, we consider the overall empirical data supporting the AOP to be MODERATE (the median call). Rank order (provided in the overall assessment Table - File:AssessmentSummaryAop-15.pdf):
Rank order of the KERs and the weight of evidence for the essentiality all point to the overall weight of evidence for this AOP as strong. Biological plausibility is strong for all KERs, with primarily moderate evidence for KER linkages and relatively few uncertainties or inconsistencies.
Known Modulating Factors
Quantitative Understanding
As described above, it is established that alkyl adducts, mutations in spermatogonia and mutations in offspring all increase with dose in a manner that is consistent with the AOP. Alkylation must exceed a threshold (determined by saturation of the relevant DNA repair pathways) before alkyl DNA lesions persist, and mutations subsequently begin to occur. However, the precise quantitative relationship has not been modeled. Existing data published in the literature could be mined to do this and thresholds for specific adduct types (i.e., estimates of how many adducts are needed to cause a mutation in a gene on average) have been published for certain cell types, which should theoretically correlate with germ cell mutagenicity for ENU and other alkylating agents.
The quantitative relationship between mutations in sperm and mutations in the offspring has not been determined and will be locus- and mutation-type specific (e.g., stronger selection against coding mutations than non-coding mutations, which will influence transmission probability); however, although many mutations will lead to embryonic loss, a large subset of mutations is expected to be heritable and viable. It is expected that quantitative understanding of this relationship will increase as advanced single cell sequencing technologies are more developed to query mutations in sperm versus offspring. For non-coding sites (e.g., transgenic reporter genes and non-coding DNA like tandem repeats), the relationship is expected to approach 1:1.
Overall, the variables that could be used to predict whether a heritable mutation is probable following exposure to an alkylating agent are the number and types of adducts per nucleotide (and knowledge of their repair efficiency). Generally, the probability of a mutation occurring is highly dependent on the type of adduct formed (mutagenicity of the adduct is based on repair efficiency and probability of error-free replication over the lesion) and abundance of the adducts, and could be modeled using existing published data.
Considerations for Potential Applications of the AOP (optional)
The information provided in this AOP will provide context for understanding how to interpret new data produced from the rodent transgene mutation assay applied to sperm (OECD TG 488) [OECD 2013], which is being increasingly applied, as well as data produced using tandem repeat mutation assays. In addition, it is envisioned that next generation sequencing technologies will enable the analysis of germ cell mutations in human populations and the eventual discovery of human germ cell mutagens. It is important to note that the regulation of chemicals that can induce heritable effects has, to date, been based heavily on extrapolation from somatic cell data. Although regulatory agencies around the world have policies in place for germ cell mutagens, risk management based on an agent that is classified as a germ cell mutagen has not yet occurred because of lack of solid evidence that these exist. This AOP demonstrates strong evidence to support the existence of male rodent germ cell mutagens, supported by data in other species (fish, flies, birds), and strongly implies that such mutagens will also affect human germ cells.
References
Allay, E., M. Veigl and S.L. Gerson (1999), "Mice over-expressing human O6 alkylguanine-DNA alkyltransferase selectively reduce O6 methylguanine mediated carcinogenic mutations to threshold levels after N-methyl-N-nitrosourea", Oncogene, 18(25): 3783-3787.
Bernstein, B.E., E. Birney, I. Dunham, E.D. Green, C. Gunter and M. Snyder (2012), "An integrated encyclopedia of DNA elements in the human genome", Nature, 489(7414): 57-74.
Boyko, A.R., S.H. Williamson, A.R. Indap, J.D. Degenhardt, R.D. Hernandez, K.E. Lohmueller, M.D. Adams, S. Schmidt, J.J. Sninsky, S.R. Sunyaev, T.J. White, R. Nielsen, A.G. Clark and C.D. Bustamante (2008), "Assessing the evolutionary impact of amino acid mutations in the human genome", PLoS Genetics, 4: e1000083.
Conrad, D.F., J.E. Keebler, M.A. DePristo, S.J. Lindsay, Y. Zhang, F. Casals, Y. Idaghdour, C.L. Hartl, C. Torroja, K.V. Garimella, M. Zilversmit, R. Cartwright, G.A. Rouleau, M. Daly, E.A. Stone, M.E. Hurles and P. Awadalla (2011) "Variation in genome-wide mutation rates within and between human families", Nature Genetics, 43(7): 712-714.
Girirajan, S., C.D. Campbell and E.E. Eichler (2010) "Human Copy Number Variation and Complex Genetic Disease", Annual Review of Genetics, 45: 203-226.
Hoischen, A., B.W. van Bon, C. Gilissen, P. Arts, B. van Lier, M. Steehouwer, P. de Vries, R. de Reuver, N. Wieskamp, G. Mortier, K. Devriendt, M.A. Amorim, N. Revencu, A. Kidd, M. Barbosa, A. Turner, J. Smith, C. Oley, A. Henderson, I.M. Hayes, E.M. Thompson, H.G. Brunner, B.B. de Vries and J.A. Veltman (2010) "De novo mutations of SETBP1 cause Schinzel-Giedion syndrome", Nat. Genet., 42(6): 483-485.
Kong, A., M.L. Frigge, G. Masson, S. Besenbacher, P. Sulem, G. Magnusson, S.A. Gudjonsson, A. Sigurdsson, A. Jonasdottir, W.S. Wong, G. Sigurdsson, G.B. Walters, S. Steinberg, H. Helgason, G. Thorleifsson, D.F. Gudbjartsson, A. Helgason, O.T. Magnusson, U. Thorsteinsdottir and K. Stefansson (2012), "Rate of de novo mutations and the importance of father's age to disease risk", Nature, 488(7412): 471-475.
Kryukov, G.V., L.A. Pennacchio and S.R. Sunyaev (2007), "Most rare missense alleles are deleterious in humans: implications for complex disease and association studies", American Journal of Human Genetics, 80(4): 727-739.
Ku, C.S., V. Vasiliou and D.N. Cooper (2012), "A new era in the discovery of de novo mutations underlying human genetic disease", Human Genomics, 12(6):27.
Lupski, J.R. (2010), "New mutations and intellectual function", Nature Genetics, 42(12): 1036-1038.
Linschooten, J.O., N. Verhofstad, K. Gutzkow, A.K. Olsen, C. Yauk, Y. Oligschläger, G. Brunborg, F.J. van Schooten and R.W. Godschalk (2013), “Paternal lifestyle as a potential source of germline mutations transmitted to offspring”, FASEB J, 27: 2873-28749. Paul C, Robaire B. 2013. Ageing of the male germ line. Nat Rev Urol 10(4):227-234.
McLaughlin, H.M., R. Sakaguchi, C. Liu, T. Igarashi, D. Pehlivan, K. Chu, R. Iyer, P. Cruz, P.F. Cherukuri, N.F. Hansen, J.C. Mullikin, Program NCS, L.G. Biesecker, T.E. Wilson, V. Ionasescu, G. Nicholson, C. Searby, K. Talbot, J.M. Vance, S. Zuchner, K. Szigeti, J.R. Lupski, Y.M. Hou, E.D. Green and A. Antonellis (2010), "Compound heterozygosity for loss-of-function lysyl-tRNA synthetase mutations in a patient with peripheral neuropathy", Am. J. Hum. Genet., 87(4): 560-566.
Morrow, E.M. (2010), "Genomic copy number variation in disorders of cognitive development", Journal of the American Academy of Child and Adolescent Psychiatry, 49(11): 1091-1104.
O'Roak, B.J., L. Vives, S. Girirajan, E. Karakoc, N. Krumm, B.P. Coe, R. Levy, A. Ko, C. Lee, J.D. Smith, E.H. Turner, I.B. Stanaway, B. Vernot, M. Malig, C. Baker, B. Reilly, J.M. Akey, E. Borenstein, M.J. Rieder, D.A. Nickerson, R. Bernier, J. Shendure and E.E. Eichler (2012), "Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations", Nature, 485(7397): 246-250.
Roach, J.C., G. Glusman, A.F. Smit, C.D. Huff, R. Hubley, P.T. Shannon, L. Rowen, K.P. Pant, N. Goodman, M. Bamshad, J. Shendure, R. Drmanac, L.B. Jorde, L. Hood, D.J. Galas (2010), "Analysis of genetic inheritance in a family quartet by whole-genome sequencing", Science, 328(5978): 636-639.
Sun, J.X., A. Helgason, G. Masson, S.S. Ebenesersdottir, H. Li, S. Mallick, S. Gnerre, N. Patterson, A. Kong, D. Reich and K. Stefansson (2012), "A direct characterization of human mutation based on microsatellites", Nature Genetics, 44(10): 1161-1165.
Thomas, A.D., G.J. Jenkins, B. Kaina, O.G. Bodger, K.H. Tomaszowski, P.D. Lewis, S.H. Doak, G.E. Johnson (2013), "Influence of DNA repair on nonlinear dose-responses for mutation", Toxicol. Sci., 132(1): 87-95.
Vissers, L.E., J. de Ligt, C. Gilissen, I. Janssen, M. Steehouwer, P. de Vries, B. van Lier, P. Arts, N. Wieskamp, M. del Rosario, B.W. van Bon, A. Hoischen, B.B. de Vries, H.G. Brunner, J.A. Veltman (2010), "A de novo paradigm for mental retardation", Nature Genetics, 42(12): 1109-1112.
Yauk C.L., Aardema, M.J., Benthem, J., Bishop, J.B., Dearfield, K.L., DeMarini, D.M., Dubrova, Y.E., Honma, M., Lupski, J.R., Marchetti, F., Meistrich, M.L., Pacchierotti, F., Stewart, J., Waters, M.D., Douglas, G.R. (2013), "Approaches for identifying germ cell mutagens: Report of the 2013 IWGT workshop on germ cell assays", "Mutation Research Genetic Toxicolology and Environmental Mutagenesis", 783: 36-54.
Xiao, W. and T. Fontanie (1995), "Expression of the human MGMT O6-methylguanine DNA methyltransferase gene in a yeast alkylation-sensitive mutant: its effects on both exogenous and endogenous DNA alkylation damage", Mutat. Res, 336(2): 133-42.